
Home
Studies
& Data Analysis
Methods
Microscope studies
Flagella experiment
Laboratory math
Blood fractionation
Gel electrophoresis
Protein gel analysis
Mitochondria
Concepts/ theory
Keeping a lab notebook
Writing research papers
Dimensions & units
Using figures (graphs)
Examples of graphs
Experimental error
Representing error
Applying statistics
Principles of microscopy
Solutions & dilutions
Protein assays
Spectrophotometry
Fractionation & centrifugation
Radioisotopes and detection
Guide to the study
Lab part 1
Lab part 2
Lab part 3
Selected methods
Organization of the Erythrocyte Membrane
Reference: Luna, EJ and AL Hitt, Cytoskeleton-plasma membrane interactions, Science 258: 955-964, 1991
The basic structure of the erythrocyte membrane cytoskeleton has been worked out, although some details remain obscure. Questions about the precise organization of the cytoskeleton and how it changes during deformation of the cell still remain. The genetic control of the multigene families of different cytoskeletal proteins remain to be worked out. Even when the last questions about the structure and function of the erythrocyte membrane are answered, the model will remain useful as a starting point in the study of other more complex cytoskeletal systems. Membrane-associated cytoskeletal protein networks are involved in the control of cell shape, attachments to other cells and to the substrate, and in organization of specialized membrane domains. Of current interest, for example, is the role of the membrane cytoskeleton in the organization of receptors and in the dynamics of receptor organization and reorganization under different conditions.
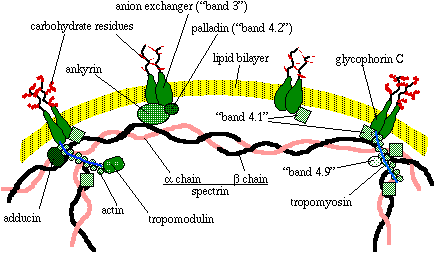
Since details of the organization of the erythrocyte cytoskeleton are still to be worked out, the cartoon presented here must be considered only a working model. Details may change and may indeed be presented differently in textbooks and review articles that you happen across.
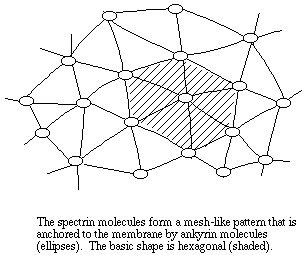
The most prominent component of the red cell membrane cytoskeleton, by molecular mass at least, is a fibrous polypeptide called spectrin. Two isoforms of spectrin, alpha (260 kDa) and beta (225 kDa), form a loosely wound helix. Two alpha-beta helixes are linked end to end to form a single tetramer, which has binding sites for several other proteins, including other spectrin molecules. The spectrin tetramers are organized into a meshwork that is fixed to the membrane by the protein ankyrin (215 kDa).
Ankyrin is itself connected to a transmembrane protein called 'band 3' or anion exchanger protein (90 to 100 kDa). The purpose of band 4.2 (palladin, 72 kDa) may be to stabilize the link between ankyrin and the anion exchanger (see the note on naming proteins by band number at the end of this document). Spectrin is also linked to a transmembrane protein called glycophorin C (25 kDa) by the protein known as 'band 4.1.' Thus the meshwork is anchored to the membrane at multiple sites.
Band 4.1 (78 kDa) stabilizes the association of spectrin with actin (subunit mass of 43 kDa), as does the protein adducin (isoforms of 100 and 105 kDa). Actin subunits actually form short microfilaments consisting of filamentous actin and tropomyosin (isoforms of 27 and 29 kD). The protein tropomodulin is also associated with filamentous actin. Band 4.9 protein, known also as dematin (48 kDa) may crosslink some of the actin microfilaments to make bundles of f (filamentous) actin.
The structure of the lipid bilayer is critical to the organization of any cytoskeletal network. For example, the transmembrane proteins require a specific environment in order to remain stable within the plasma membrane. Types of phospholipids vary, and not every combination will work. Consider, for example, that ankyrin has an attached fatty acid called palmitate, which may fit into the lipid bilayer itself, further stabilizing its attachment to the cell membrane.
One means of regulating protein function is by the addition of one or more phosphate groups to a protein by enzymes called phosphoprotein phosphatases. Addition of a phosphate group is called phosphorylation, and removal is called dephosphorylation. Phosphorylation of major proteins such as ankrin and band 4.1 and 4.9 proteins can weaken the rigidity of the cytoskeleton by reducing the binding affinity of these components. Thus rigidity of the cytoskeleton is under some control. Precisely how that control is exercised is of current interest to cell biologists.
Why name proteins by band number?
Some of the earliest work on membrane cytoskeletal proteins involved electrophoresis of denatured membrane proteins on SDS gels (SDS-PAGE). Such analysis revealed the presence of specific protein bands that were later associated with structures that were apparent in electron micrographs, that had similarities to familiar cytoskeletal proteins, etc. Since the proteins that were identified were unknown, they were identified by band number.Later, it was discovered that some of the numbered proteins had already been identified and characterized in different tissues. Since so much of the literature already referred to proteins by number, some of the 'band' names stuck. You can probably guess what kind of confusion would take place if the same protein had multiple names, or if the names kept changing throughout the literature. When new macromolecules are discovered independently they are often named separately before any similiarities are noticed. It is necessary, then, to stick with the most well-known names even if they are not terribly informative or otherwise appropriate.
Visitors: to ensure that your message is not mistaken for SPAM, please include the acronym "Bios211" in the subject line of e-mail communications
Created by David R. Caprette (caprette@rice.edu), Rice University 17 May 96
Updated 30 Jul 12
