
Home
Studies
& Data Analysis
Methods
Microscope studies
Flagella experiment
Laboratory math
Blood fractionation
Gel electrophoresis
Protein gel analysis
Mitochondria
Concepts/ theory
Keeping a lab notebook
Writing research papers
Dimensions & units
Using figures (graphs)
Examples of graphs
Experimental error
Representing error
Applying statistics
Principles of microscopy
Solutions & dilutions
Protein assays
Spectrophotometry
Fractionation & centrifugation
Radioisotopes and detection
Guide to the study
Lab part 1
Lab part 2
Lab part 3
Selected methods
Example of a Standard Curve for Molecular Mass
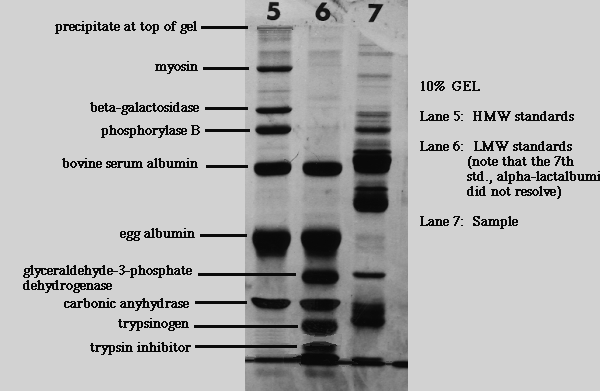
Protein standards for gels are purified polypeptides with relative mobilities that correspond closely to their true molecular mass. Suppliers of chemicals for electrophoresis such as SIGMA (St. Louis, MO) or Bio-Rad (Hercules, CA) provide ready-made molecular mass standards. SIGMA sells standards for calibrating SDS gels with a tris based buffer system (Laemmli gels).
Inserts with commercial standards, older literature, and even my old web pages may refer to molecular weight (MW) rather than molecular mass. MW is the same as relative molecular mass (Mr) but differs from molecular mass in that it is a unitless quantity. You will likely find MW data that are reported in Daltons or kiloDaltons although such usage is not correct. A number representing MW is identical to the corresponding molecular mass in Daltons, thus the terms can be used interchangeably as long as the units are dropped from quantities that are represented as MW.
SIGMA Standard Mixture for Molecular Weights 30,000-200,000 (SDS6H2)
- myosin from porcine muscle, 200,000
- beta-galactosidase, from Escherichia coli, 116,000
- phosphorylase B, from rabbit muscle 97,400
- albumin, bovine 66,000
- albumin, from chicken egg white 45,000
- carbonic anhydrase from bovine erythrocytes, 29,000
SIGMA Dalton Mark VII-L Standard Mixture, MW range 14,000-70,000 (SDS-7)
- albumin, bovine 66,000
- albumin, from chicken egg white 45,000
- glyceraldehyde-3-phosphate dehydrogenase, from rabbit muscle 36,000
- carbonic anhydrase from bovine erythrocytes, 29,000
- trypsinogen, from bovine pancreas 24,000
- trypsin inhibitor, soybean, 20,100
- alpha-lactalbumin, bovine milk 14,200
Note that the sources of proteins are varied. You won't find all of them in any one protein fraction, in fact you aren't likely to find any of them, depending on the fraction you are studying. They are used to calibrate gels, not as indicators of what types of proteins are present. Keep in mind that many very different proteins have similar molecular weights. The patterns given by standard mixes become recognizable with experience.
A typical plot of the molecular weights of standards versus their relative mobilities is shown below, using a log scale for the molecular weights. Be very careful with curve fits, keeping in mind that the scale is logarithmic. You don't want to be in error near the top of the gel. It may be better to simply interpolate results (connect the data points).
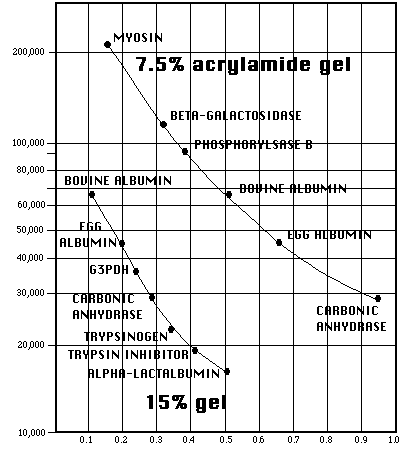
Relative mobility for a given polypeptide will vary with gel density (%T). Curves will always shift when percent acrylamide changes.
With some gels of high percent acrylamide, the dye front will not be evident, since proteins small enough to run with the same mobility as the bromphenol blue dye may not be present. Unless the position of the dye front was marked before staining, a true relative mobility cannot be determined. As long as a gel is calibrated using internal standards, MW estimates can be obtained by representing relative migration distance using an arbitrary reference point such as the bottom of the gel. Use of a true relative mobility allows one to use the same standard curve for any gel of the exact same composition regardless of dimensions or position of the dye front when electrophoresis is terminated. Error in MW estimates is compounded with the use of an arbitrary reference point when the original dye front was curved or otherwise distorted.
Visitors: to ensure that your message is not mistaken for SPAM, please include the acronym "Bios211" in the subject line of e-mail communications
Created by David R. Caprette (caprette@rice.edu), Rice University 17 May 96
Updated 25 May 05
