EVOLUTIONARY GENOMIC ANALYSES OF SPECIES DIVERGENCE
We are interested in the origin, population genetics, and molecular evolution of genomic novelties, such as gene duplications or gene expression changes that uniquely distinguish closely related species.
 Moreover, we are interested in understanding why the genomes of closely related species remain porous for some parts of the genome but impermeable for others.
Moreover, we are interested in understanding why the genomes of closely related species remain porous for some parts of the genome but impermeable for others.
Are there any generalities that could be described for genomic regions that can or cannot be exchanged between species? What are the traits these genes encode? Are these genes and traits adaptive and can the signature of selection be detected at the level of the sequence?
Closely related species are particularly useful to study the evolution of the genome in the context of species differentiation and the processes promoting it. We are particularly interested in studies of genome divergence of genetically equidistant pairs of species that are allopatric and sympatric/parapatric. Models of sequence divergence and speciation make rather dramatically different predictions regarding the patterns of genome divergence between species under these geographic scenarios.
Introgression of the vitamin K cycle from Mus spretus into the common house mouse
It is rarely possible to observe the natural transfer of genes underlying an adaptive trait between species and to follow the outcome. We have discovered such adaptive introgressive hybridization between two species of mice.
 The process has introduced a highly divergent version of vkorc1 of M. spretus into the European population of common house mice, where it confers warfarin resistance and has spread far beyond the hybridization area in SW Europe into Germany.
The process has introduced a highly divergent version of vkorc1 of M. spretus into the European population of common house mice, where it confers warfarin resistance and has spread far beyond the hybridization area in SW Europe into Germany.
One particularly interesting question we are pursuing refers to the effect the introduction of such a divergent copy of vkorc1, which has evolved at the high rate of all sequenced genes of M. spretus (Ka/Ks>1.8) and at functionally crucial positions, has on the population genetics and molecular evolution of genes that depend on it to maintain vitamin K recycling efficiency, blood coagulation, and other processes. The gene has been highly constraint during its evolution from invertebrates to human.
We distinguish between two scenarios that could compensate for the introgression of such a divergent gene. One such scenario involves the co-introgressionn of genes that interact with vkorc1. Alternatively, or in addition, compensatory mutations in interacting genes may enable the gene to function in the genomic background of house mice. We are setting up in vitro studies that quantify the vitamin K cycle kinetics if alleles at various interacting genes are expressed (by co-transfected cells) that stem from house mice only or are combined with alleles of M. spretus.


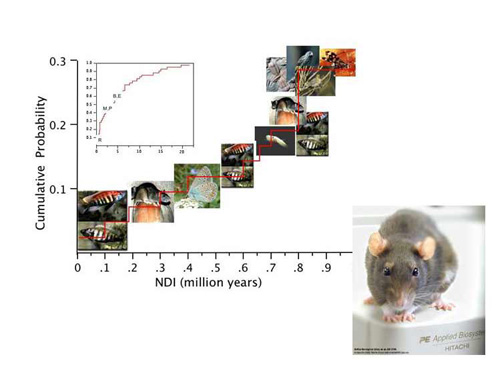
Genomic differentiation in Rattus
In the long term we wish to explore the research opportunities offered by the recent origin, rapid rate of speciation and niche diversification, and the now vast geographic range of more than 60 closely related Rattus species in SE Asia and Australia.
To study the genome evolution in this group we are using the published rat genome sequence and other genomic resources to examine how biogeography and natural selection have interacted to generate such diversity at the taxonomic and genomic level in less than 2-3 million years. Next-generation sequencing technologies are used to collect genome divergence data in Rattus.
Inferences of the genetics of primate speciation
|
Macaca fascicularis, one species of macaque that forms hybrids with the rhesus monkey, M. mulatta, in SE Asia |
While much of our work is concerned with rats and mice we are also exploring a primate system. Our work has further identified an area in SE Asia where two species of monkey, Macaca mulatta and M. fascicularis, form hybrids. We have found preliminary evidence that not all genes are equally likely to be transferred between the two species. What makes these genes that do seem to be less likely to be transferred between the species special? A non-human primate system could be of great value because it may yield insights into the similarities and differences in primate and rodent genome differentiation and speciation. For example, considering the extraordinary evolution of the primate brain, social systems, and behaviors; could it be that genes underlying cognitive processes have played a crucial role in primate speciation? |
